By: Laura Tibbs-Cortes | 01/30/2025 ORISE Postdoctoral Fellow, USDA-ARS Crop Genome Informatics Laboratory, Ames, IA
In maize and other crops, important traits are often complex, affected by genetics, the environment, and their interaction. In addition, different crop varieties exhibit varying degrees of phenotypic plasticity, in which a given genotype displays different phenotype values in different environments. Identifying genes influencing traits as well as their plasticity can help plant breeders to develop successful new varieties, especially in the face of changing climate.
In a recently published project, SCINet/AI-COE Postdoctoral Fellow Laura Tibbs-Cortes, working in the lab of Dr. Xianran Li, in collaboration with Dr. Jianming Yu at Iowa State University, worked to systematically identify environmental and genetic determinants of a wide variety of traits in maize. This project leveraged open science data in which 19 traits were measured in each of 5,000 maize varieties across 11 environments. From these data, the CERIS-JGRA framework (Li et al. 2021, https://doi.org/10.1016/j.molp.2021.03.010) was used to identify environmental indices based on temperature, precipitation, day length, or combinations of these variables that were most strongly correlated with variation in the traits of interest. The environmental indices themselves were biologically relevant (Fig. 1) and robust to sub-sampling. By using these environmental factors as regressors, genotype-specific models of phenotypic plasticity were built, with the slopes quantifying genotype-specific trait plasticity. These models enabled accurate performance prediction.
Genetic loci affecting trait means and plasticities were identified using genome-wide association studies (GWAS) using more than 20 million markers recently generated by high-density sequencing efforts. Leveraging these high-density genomic data relied on high-performance computing resources provided by SCINet and enabled this project to uncover additional patterns in phenotypic plasticity that were not found in previous analyses using fewer markers.
Model parameters from CERIS-JGRA (slope and intercept) were used as input phenotypes for GWAS. This approach successfully identified loci significantly associated with all 19 traits; the generated candidate gene list is publicly available and represents a community resource for ongoing study of maize plasticity. In addition, studying a wide variety of traits enabled the creation of conceptual figures to visualize the relationships among important parameters under current hypotheses of phenotypic plasticity. The results of this project provide insight not only into the genetic architecture of phenotypic plasticity in maize but also an updated perspective on the ongoing debate between alternative models of phenotypic plasticity.
This project featured collaboration across multiple institutions, including two USDA units: the Wheat Health, Genetics, and Quality Research Unit in Pullman, Washington and the Corn Insects and Crop Genetics Research Unit in Ames, Iowa.
Results have been uploaded as a track on the genome browser at MaizeGDB, a USDA-funded community resource for maize genetics and genomics (https://jbrowse.maizegdb.org/?data=B73&tracks=Tibbs-Cortes2024%2Cgene_models_official).
For more details, check out our open-access publication in Genome Research at https://www.genome.org/cgi/doi/10.1101/gr.279027.124.
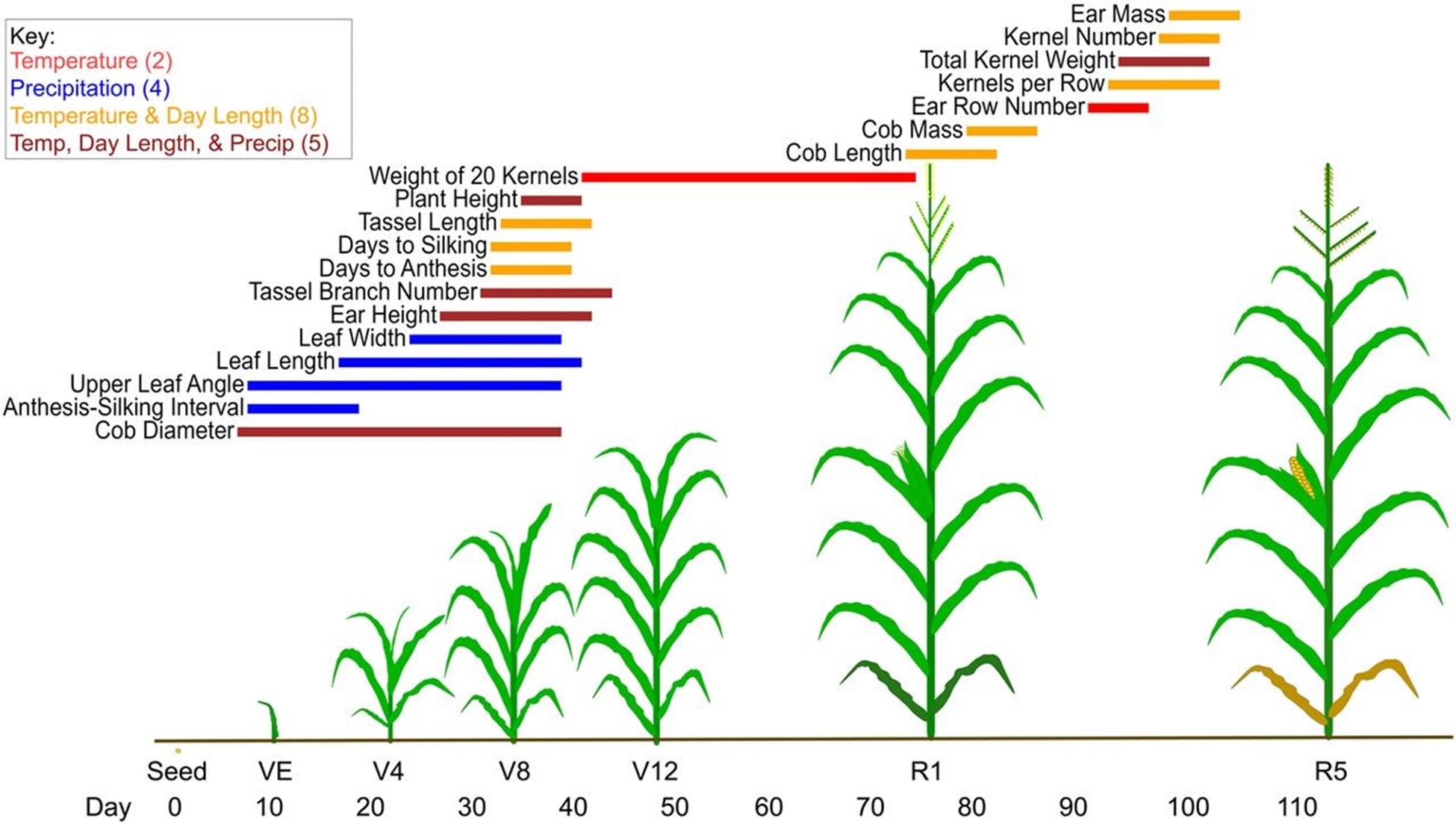
Figure 1: Environmental indices identified for the 19 traits, shown with an example of maize development. For each trait, a line segment represents the environmental index identified by CERIS-JGRA, with color denoting the environmental variable(s) used and extent on the x axis the time window. Environmental indices show biological relevance, with a clear transition from the vegetative to the reproductive stage visible.

